Cardiolund ECG Parser
Product overview
The Cardiolund ECG Parser is a medical software for automated rhythm analysis and categorization of 1-12 lead ECGs.
The software provides automated ECG signal analysis with interval measurements and categorization approaching the level of a professional trained in cardiology, including:
- Robust estimation of standard ECG parameters in ECG signals.
- Detection and classification of beats and rhythm.
- Categorization of ECGs into clinically useful groups (12 categories: Poor Quality, No Rhythm Deviation, Irregular Sequence, etc.), to enable prioritized presentation of signals in a user interface.
The ECG Parser provides analysis results as a machine-readable data structure. The software is intended for integration with medical systems, or mobile device user interfaces. End users, such as clinical staff and patients, interact with the ECG Parser through the interface of the medical system and/or the device UI.
The product features are described in more detail in the following.
If you need more information on the EGG Parser product, or want to enquire about purchasing options, please contact us.
Main features
The ECG Parser software provides a thorough rhythm analysis of ECG recordings with 1-12 leads and durations of ten seconds up to days or weeks. The ECG Parser
- Identifies beats and groups them into beat classes with different morphologies.
- Analyses the beat sequence in detail providing markers for a wide range of events.
- Calculation of standard measures such as Heart rate, RR variability, and the RR corrected QT interval (QTc) is included, as well as R tag and QT interval markers for presentation in a user interface.
- Performs robust interval measurements based on averaged beats, and
- Categorizes signal segments into one of 12 categories for efficient identification of important cases in large databases or important segments in long recordings.
The parsing process produces a wealth of metadata that describe the contents of the recording, including the pre-processed ECG signal, averaged beats and associated measures, beat markers and beat classification, and markers and counters for short/long beats, SVES/VES, AV block II and longer pauses, faster or slower sequences, bigemini/trigemini as well as segments of irregularity. The metadata can be used for generating custom reports, creating plots of calculation results, and for display in a user interface.
The output of the ECG Parser is provided in a machine-readable format, and it is therefore well-prepared for subsequent statistical analysis, and for applying machine learning and artificial intelligence techniques.
The ECG Parser is meant for integration with other software, and therefore it does not provide a graphical user interface, nor does it contain software for data capture and storage. The software can be customized and extended to provide different types of analysis reports, but it can also serve as an analysis platform supporting the customer device’s own user interface. Analysis reporting is created according to customer specific requirements, e.g., design, layout, localization to specific languages, etc.
Categorisation, tags and markers
The ECG Parser categorizes each signal (with length ranging from 10 to 60 s) into one of the following 12 categories:
| # | Category | Deviation |
|---|---|---|
| 0. | Poor Quality | Difficult to follow the beat sequence |
| 1. | No Rhythm Deviation | Only insignificant irregular beats present |
| 2. | Irregular Sequence | Unexplained irregularities which may be atrial fibrillation (AF) or paroxysmal AF with or without P waves |
| 3. | Pause/AVblockII | AV block 2 or beats longer than 2.2 s |
| 4. | Fast Regular | Fast rhythm without wide QRS complexes, RR interval shorter than 600 ms |
| 5. | Fast Regular and Wide QRS | Fast rhythm with wide QRS complexes, RR interval shorter than 600 ms and QRS duration more than 135 ms |
| 6. | Fast/Slow Sequences | Shorter sequences of faster or slower beats |
| 7. | Bigemini | Bigemini patterns with P waves detected |
| 8. | Trigemini | Trigemini patterns with P waves detected |
| 9. | Wide QRS | QRS duration more than 135 ms in general, but not fast |
| 10. | >5 SVES | More than 5 supraventricular extrasystole detected |
| 11. | >5 VES | More than 5 ventricular extrasystole detected, not all wide beats |
The analysis results of a signal also contain tags, filtered versions of the signal, and a detailed set of descriptors and markers for identified events in the signal.
Categories and tags can be used for prioritizing resources during screening, since it allows the clinician to focus on the important cases, i.e., recordings classified as having a rhythm deviation. The clinician can thereby dismiss most of the recordings, namely all those classified as having poor quality, or as having no rhythm deviation.
The metadata provided with the analysis results can be used to annotate the signal when displaying it in a graphical user interface, as shown in the following figures.
Annotation of long beats:
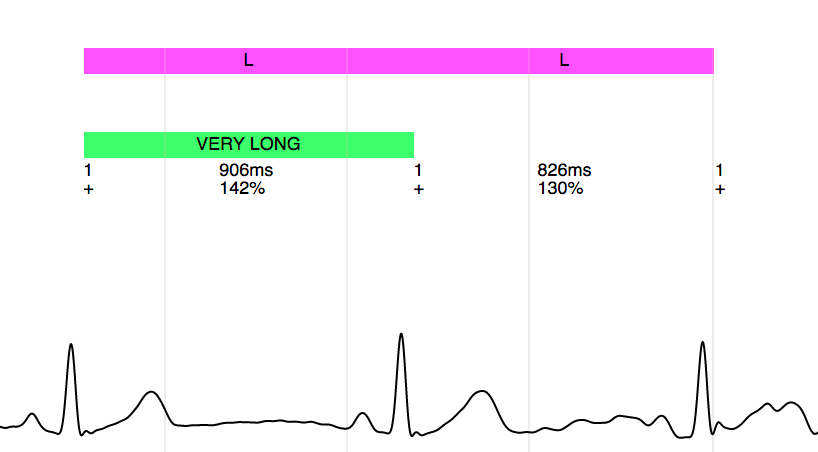
Annotation of a SVES (supraventricular extrasystole):
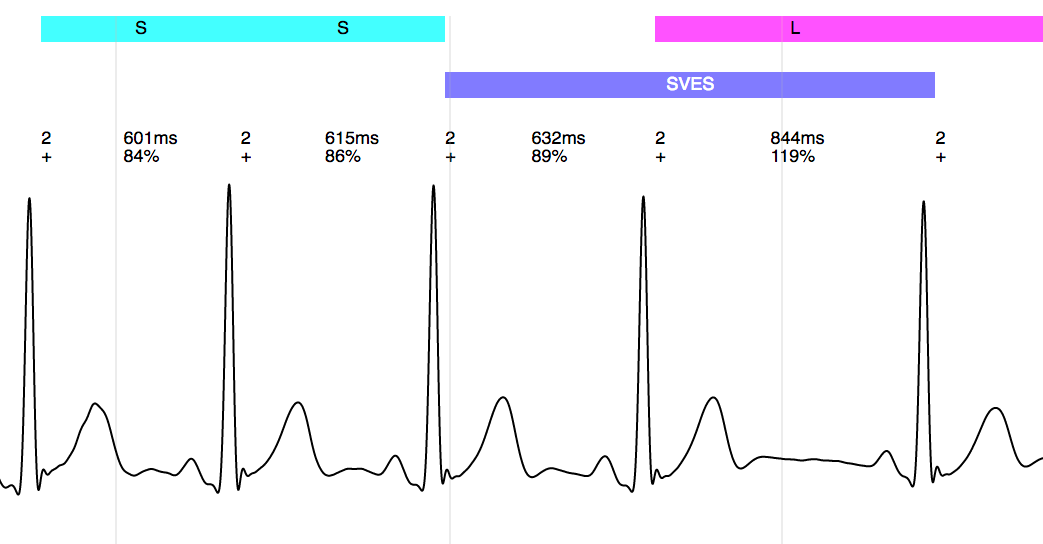
Clinical applications
One important clinical application of the ECG Parser is screening for atrial fibrillation (AF). When such screening is based on simplified home-based ECG measurements, the number of measurements is often very large, making it practically impossible to manually review all ECG recordings. The ECG Parser is well-matched to this application and fulfils an urgent need for software which can parse the signals and select a smaller group requiring further review.
Integrating the ECG Parser in a screening system for diagnosis of AF only requires the manufacturer to choose a subset of the above categories as the “AF detector”. Different selections of categories are associated with different sensitivity and specificity. Cardiolund has clinical data available that shows how these two performance measures vary with choice of categories. When selecting a certain subset of categories as AF detector, it is important to notice that, in clinical practice, AF is often grouped with other arrhythmias, in particular atrial flutter.
Other important clinical applications include analysis of Holter recordings, detection of serial changes in event recordings, the identification of common properties in signals from patient with the same end-state, the identification of patients responding to treatment, and the search for important events in the data records.
Clinical performance
The ECG Parser has been thoroughly validated in published clinical studies and in performance evaluations against international standards. It is validated that the ECG Parser:
- Performs interval measurements with correct beat detection with high accuracy.
- Performs equally or better than reference systems available on the market in identifying AF from single-lead event recordings, when comparing the data available in published clinical studies.
- Classifies each segment into 12 categories representing the ECG signal content.
- Detects and classifies (supraventricular and ventricular) ectopic beats and deviating rhythm, including AF and VF.
Only heart rate (HR), interval measurements, categorization, beat and rhythm classification are validated in the ECG Parser output. Other output parameters may be used for indication only.
The STROKESTOP study
The ECG Parser has been validated in a clinical study reported in the paper:
Emma Svennberg, Martin Stridh, Johan Engdahl, Faris Al-Khalili, Leif Friberg, Viveka Frykman, Mårten Rosenqvist; Safe automatic one-lead electrocardiogram analysis in screening for atrial fibrillation. Europace 2016 euw286.
doi: 10.1093/europace/euw286
The paper describes the results of the STROKESTOP screening study conducted on 3209 persons randomly selected among a large group of 78 or 79 year olds. The data set contains a collection of 80149 thumb-ECG recordings each 30s long, which were recorded using single-lead (lead-I) ECG recording devices created and marketed by Zenicor Medical Systems. The recordings were made twice daily over a period of 14 days, by the participants themselves in the participants’ own home and without any supervision or assistance. The performance of the ECG Parser algorithm for AF screening is evaluated on the data set against a gold standard of manual review. The reported performance results are listed in the summary table, in the main report.
In the referenced paper, the ECG Parser category set of categories 2-8 and 10-11 are used as AF diagnostic indicator. The study concludes that no AF case is missed in the screening process using these ECGP categories as AF diagnostic indicator.
Performance results for other choices of category sets for AF indication can be provided by Cardiolund on request. Cardiolund has also performed an additional performance assessment of the ECG Parser regarding categorization of recordings into the 12 categories based on a sub-set of the STROKESTOP data set, and on additional clinical assessment.
Clinical performance summary
The clinical performance of the ECG Parser depends to a large extent on the signal quality of the ECG recordings and may therefore vary between different systems.
The ECG Parser has the following documented clinical performance:
-
The AF screening performance has been evaluated using the 80k signals in the STROKESTOP study measured using a handheld Zenicor ECG device. For this dataset, an AF sensitivity of 100% on a signal/patient level is achieved with a specificity of 88%.
-
The categorization performance has been evaluated using 4k signals measured using a handheld Zenicor ECG device. For this dataset, the categorization has shown to be indicative of important clinical characteristics of the signal. The performance is measured in terms of specificity exceeding 97% for all rhythm deviation categories.
-
The beat detection and interval measurement (including QT-interval measurements) performance has been evaluated using the CSE, CST, and the Physionet QT, MIT, AHA, NST databases, and using requirements from international standards IEC 60601-2-25 and IEC 60601-2-47.
-
Detection and classification of ectopic beats and deviating rhythm, including AF and VF, performance has been evaluated using the Physionet MIT, AHA, NST databases, and using requirements from the international standard IEC 60601-2-47.
Integrating with a medical device
The ECG Parser can be integrated with custom software and ECG recording hardware to form a medical device system that provides automated interpretation of 1-12 lead ECGs. The ECG Parser can handle varying levels of signal quality, ranging from poor-quality single-lead ECGs acquired with mobile devices, to high-quality 12-lead recordings.
The ECG Parser is is not a complete diagnostic software, neither is it intended for real-time applications, nor for interpretation of all life-threatening cardiac diseases.
An ECG system that integrates the ECG Parser is typically classed as a Class IIa medical device on the EU MDR classification scale, and as Class B (Non-serious injury possible) on EN IEC 62304 classification scale.
With a Class IIa system classification, the analysis results of the ECG Parser can be used for the following post-processing applications:
- ECG categorization
- ECG prioritization
- ECG screening, in particular screening for atrial fibrillation (AF)
- Holter ECG reporting
With a Class I system classification, the analysis results ECG Parser can be used for the following (diagnostics or treatment evaluation) applications:
- ECG decision support
- ECG pre-processing for subsequent aggregation
- ECG-related workflow improvement in user interfaces
- ECG drug response analysis
- ECG database search applications
There are also many possible applications of the software that does not invoke medical device legal requirements, including the following (plotting, storage, research and non-clinical) applications:
- ECG research
- ECG visualization
- ECG storage
- ECG-related biometric applications
The ECG Parser is only intended for analysis of ECG signals recorded on humans. The software may be used for analyzing ECG signals recorded on children when the results are first reviewed by trained professionals. The software has also been used successfully for analysis of ECG signals recorded on animals.
How the ECG Parser works
A main challenge in analysis of a lower quality ECG recording is to separate irregular beat patterns from disturbance (noise) patterns. The algorithm in the ECGP software is designed and tuned for:
- Finding irregularities in beat sequences in the presence of the disturbance patterns which are associated with lower quality ECG measurements.
- Not missing any important cases and, at the same time, significantly reduce the required review effort by a trained cardiologist.
The main principle of the algorithm contained in the ECG Parser is to detect all beats, events, and disturbances, and evaluate the most common beat morphologies and the rhythm patterns of those. The analysis is divided into the following steps:
- Segmentation of longer signals to allow for parallel processing in subsequent steps.
- Re-sampling to adjust the signal to the internal sampling rate of the algorithm.
- Preprocessing to remove disturbances that may disturb the beat detector.
- Stim detection and removal to remove pacemaker spikes that may disturb the beat detector.
- Beat detection to locate and identify all individual beats and group them into beat classes based on morphology.
- Average beat analysis and interval measurements is where all intervals (standard ECG measures and other beat descriptors) are calculated.
- P wave analysis to identify beats where P waves are present which supports the following rhythm analysis and categorization steps.
- Rhythm analysis to identify and describe rhythm patterns and individual events, such as ectopic beats and pauses.
- Categorization and tagging are performed in the final step where all different features of the signal are summarized into a decision on which category the signal belongs to (Poor Quality, No Rhythm Deviation, Irregular Sequence, etc.).
The following diagram provides an overview of the information flow through the steps of the algorithm.
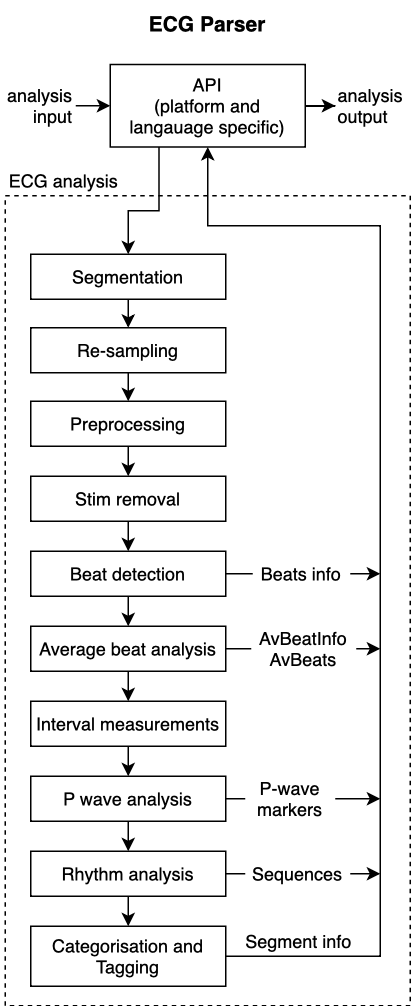
Integration of the ECG Parser with a medical device system is done on the software API level.
Detailed technical specifications are availble on request. If you want more information, or have any questions, please contact us.